Single-Cell RNA Sequencing for Plant Research
The potential of scRNA-seq technology is immense, offering insights into the intricate mechanisms of gene regulation and the identification of distinct cell types and their functions. The potential of scRNA-seq technology is immense, offering insights into the intricate mechanisms of gene regulation and the identification of distinct cell types and their functions.
CD Genomics harnessing the power of single-cell genomics platform, enables a deeper understanding of plant developmental processes and their responses to various stimuli.
Overview
It has a wide range of applications in plant research. Compared with traditional bulk RNA-seq, single-cell RNA sequencing (scRNA-seq) plant developmental processes encompass multiple regulatory factors as well as significant heterogeneity among different cells, and single-cell transcriptomics provides a breakthrough approach to delve into the unique gene expression profiles of individual cells. The ability to reveal new and rare plant cell types with precision at the individual cell level also enriches our understanding of cell developmental trajectories and cellular functions.
Currently, plant single-cell transcriptome studies are mainly carried out in species such as Arabidopsis, rice, maize, tomato and poplar. Among them, the model plant Arabidopsis thaliana is a popular choice for single-cell studies. Samples typically include a range of plant organs, including roots, stems, leaves, flowers, pollen, spermatozoa, and seed endosperm.
Due to the complex composition of plant cell walls, plant single-cell transcriptome studies fall into two main categories: protoplast RNA sequencing (scRNA-seq) and cytoplasmic transcriptome sequencing (snRNA-seq). Protoplast-based methods have been the main focus of research efforts.
The basic technical steps in our plant single-cell transcriptome analysis include tissue dissociation, protoplast preparation, single-cell capture, library construction, sequencing, and in-depth data analysis.
At CD Genomics, we integrate a variety of sequencing platforms to provide comprehensive sequencing services for spatial transcriptome, single-cell transcriptome, general transcriptome, and epigenetic sequencing (such as scATAC-seq/snATAC-seq). For more information, please feel free to consult with our dedicated technical team.
Features
| Mapping | Developmental Differentiation | Mechanism Research | Environmental Response |
|---|---|---|---|
| The cellular makeup of a species' tissues is meticulously structured to unveil its cellular diversity, facilitating the discovery of novel cell types. | Integrating mapping techniques with time series analysis offers a comprehensive approach to unraveling the developmental path of specific tissues. | Delving into the intricacies of gene and transcription factor functionality, we aim to elucidate the underlying mechanisms driving distinct biological activities, such as the regulation of stomatal aperture. | Our service endeavors involve an in-depth exploration of cellular stress response mechanisms, pinpointing the specific responsive cells and the associated genes. |
Project Workflow

1. Sample Preparation
Tissue dissociation, protoplasmic preparation, single cell capture.

2. Library Preparation

3. Sequencing
Illumina HiSeq;
PE50/75/100/150;
>10G clean data.

4. Data Analysis & Delivery
Visualize and preprocess results, and perform custom bioinformatics analysis.
Bioinformatic Analysis
Dimensionality Reduction and Clustering: Complex data is simplified to reveal cellular diversity and organization within plant tissues.
Cellular Annotation: Different cell types are identified and labeled to understand the makeup of plant samples.
Subpopulation Segmentation: Distinct cell groups within tissues are uncovered, providing a finer-grained view of cellular diversity.
Cell Developmental Trajectories: Cell development paths are traced, shedding light on growth and differentiation.
Identification of New or Rare Cellular Taxa: Previously unknown or rare cell types are discovered.
(Novel) Marker Gene Mining: A novel approach is introduced to find cell type-specific marker genes.
Time-Series Analysis: How cells change over time is examined, enhancing our understanding of plant development.
Transcription Factor-Gene Regulatory Networks: Gene regulation within different cell types is explored.
Cell Communication Analysis: The interactions between plant cells are studied, revealing communication networks.
Sample Requirements
Sample Collection
- Sample Selection: For optimal results, select young and tender plant parts for sampling. These parts are recommended due to their higher cellular activity and sensitivity to experimental conditions.
- Sample Size: Ensure that each sample weighs at least 1-2 grams for a single experiment. This ensures an adequate amount of material for analysis.
- Sample Replication: It is advisable to send 3 copies of each sample to increase the reliability of your experiment's results. This redundancy helps mitigate potential errors and provides more comprehensive data.
Sample Types
- Roots: Since roots typically have lower cellular content, it's essential to provide a more substantial sample for accurate testing.
- Leaves: Focus on sending young leaves, as they contain more chloroplasts and larger cells, making them ideal for testing.
- Fruit: Given that most fruit cells are relatively large, standard sampling practices apply.
- Stem: Stem samples require normal sample delivery.
- Flower: Flowers, especially pollen cells with thick walls, necessitate specific sampling for testing due to their complex structure.
Plant Protoplast Preparation
- Cell Wall Removal: The primary method for cell wall removal is enzymatic, employing a mixture of cellulase and pectinase. This process ensures the integrity of the protoplasts while removing the cell walls effectively.
- Quality Assessment: After cell wall removal, quality control measures are vital. Protoplast activity, concentration, and impurity levels are evaluated through manual microscopy to ensure the integrity of your samples.
Custom Requirements
For any specific or unique requirements beyond these general guidelines, please contact our technical team. We are here to assist you in tailoring our services to meet your individual needs and research goals.
Deliverable
FastQ, BAM, coverage summary, QC report, custom bioinformatics analysis.
Demo Results
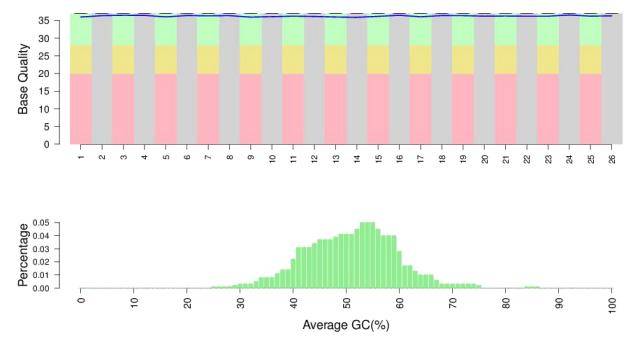 Data quality control
Data quality control
 Cell filtration and statistics
Cell filtration and statistics
 Dimensionality reduction and clustering
Dimensionality reduction and clustering
 Identification of cell subpopulations
Identification of cell subpopulations
 Differentially expressed gene analysis
Differentially expressed gene analysis
 KEGG pathway functional analysis
KEGG pathway functional analysis
 Pseudotime trajectory analysis
Pseudotime trajectory analysis
Case Studies
- Single-Cell Transcriptome Resolves Cell-Specific Stress Tolerance Gene Functions
- Single-Cell Transcriptome Explores Developmental Differentiation Trajectories Under the Influence of Stress
- Single-Cell Transcriptome Constructs Stress-Responsive Regulatory Networks
FAQ
- My samples do not have a reference genome, can I do plant single-cell transcriptome?
- Which parts of plants are generally used to prepare protoplasts?